gc3libs.optimizer.dif_evolution¶
This module implements a global optimization algorithm called Differential Evolution.
Consider the following optimization problem: 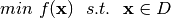 , where
, where 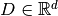 and
and 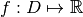 . Class
. Class DifferentialEvolutionAlgorithm solves this
optimization problem using the differential evolution algorithm. No further
assumptions on the function 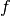 are needed. Thus it can be non-convex,
noisy etc.
are needed. Thus it can be non-convex,
noisy etc.
The domain 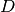 is implicitly specified by passing the function
is implicitly specified by passing the function
filtern_fn() to DifferentialEvolutionAlgorithm.
Some information related to Differential Evolution can be found in the following papers:
- Tvrdik 2008: http://www.proceedings2008.imcsit.org/pliks/95.pdf
- Fleetwood: http://www.maths.uq.edu.au/MASCOS/Multi-Agent04/Fleetwood.pdf
- Piyasatian: http://www-personal.une.edu.au/~jvanderw/DE_1.pdf
evolve_fn() is an adaptation of the following MATLAB code:
http://www.icsi.berkeley.edu/~storn/DeMat.zip hosted on http://www.icsi.berkeley.edu/~storn/code.html#deb1.
-
class
gc3libs.optimizer.dif_evolution.DifferentialEvolutionAlgorithm(initial_pop, de_strategy='DE_rand', de_step_size=0.85, prob_crossover=1.0, exp_cross=False, itermax=100, dx_conv_crit=None, y_conv_crit=None, in_domain=None, seed=None, logger=None, after_update_opt_state=[])¶ Differential Evolution Algorithm class.
DifferentialEvolutionAlgorithmexplicitly allows for an another process to control the optimization. Driver classes can be found ingc3libs.optimizer.drivers.py.Parameters: - initial_pop (list) – Initial population for the optimization.
- de_strategy (str) – e.g. DE_rand_either_or_algorithm. Allowed are:
- de_step_size (float) – Differential Evolution step size.
- prob_crossover (float) – Probability new population draws will replace old members.
- exp_cross (bool) – Set True to use exponential crossover.
- itermax (int) – Maximum # of iterations.
- dx_conv_crit (float) – Abort optimization if all population members are within a certain distance to each other.
- y_conv_crit (float) – Declare convergence when the target function is below a y_conv_crit.
- in_domain (fun) – Optional function that implements nonlinear constraints.
- seed (float) – Seed to initialize NumPy’s random number generator.
- logger (obj) – Configured logger to use.
- after_update_opt_state (list) – Functions that are called at the end of
DifferentialEvolutionAlgorithm.after_update_opt_state(). Use this list to provide problem-specific printing and plotting routines. Examples can be found ingc3libs.optimizer.extra.
The de_strategy value must be chosen from the dif_evolution.strategies enumeration. Allowed values are (description of the strategies taken from http://www.icsi.berkeley.edu/~storn/DeMat.zip):
'DE_rand': The classical version of DE.'DE_local_to_best': A version which has been used by quite a number ofscientists. Attempts a balance between robustness # and fast convergence.
'DE_best_with_jitter': Taylored for small population sizes and fastconvergence. Dimensionality should not be too high.
'DE_rand_with_per_vector_dither': Classical DE with dither to become even more robust.'DE_rand_with_per_generation_dither': Classical DE with dither to become even more robust.Choosing de_step_size = 0.3 is a good start here.
'DE_rand_either_or_algorithm': Alternates between differential mutation and three-point- recombination.
-
evolve()¶ Generates a new population fullfilling in_domain.
Return type: list of population members
-
static
evolve_fn(population, prob_crossover, de_step_size, dim, best_iter, de_strategy, exp_cross)¶ Return new population, evolved according to de_strategy.
Parameters: - population – Population generating offspring from.
- prob_crossover – Probability new population draws will replace old members.
- de_step_size – Differential Evolution step size.
- dim – Dimension of each population member.
- best_iter – Best population member of the current population.
- de_strategy – Differential Evolution strategy. See
DifferentialEvolutionAlgorithm. - bool (exp_cross) – Set True to use exponential crossover.
-
select(new_pop, new_vals)¶ Perform a one-on-one battle by index, keeping the member with lowest corresponding value.